SPANDx Genomic Profiling: Verifying LTtr and K331A RNASeq Mutations
gene_x 0 like s 1371 view s
Tags: plot, bash, tool
-
Set up the directory for raw data.
#Replace "p600" with "control", "p602" with "LT", "p605" with "LTtr", and "p783" with "K331A". Please note that the RNAseq data from the LT_K331A_d8 replicates were sequenced alongside the ChIPseq batch. ln -s /home/jhuang/DATA/Data_Denise_LT_DNA_Bindung/Raw_Data_ChIPseq/230306_NB501882_0417_AHMVHHBGXN/2023_022_nf_denise/nf857/1_NHDF_Donor_1_p783_S1_R1_001.fastq.gz LT_K331A_d8_DonorI.fastq.gz ln -s /home/jhuang/DATA/Data_Denise_LT_DNA_Bindung/Raw_Data_ChIPseq/230306_NB501882_0417_AHMVHHBGXN/2023_022_nf_denise/nf858/2_NHDF_Donor_2_p783_S2_R1_001.fastq.gz LT_K331A_d8_DonorII.fastq.gz mv V_8_2_4_p600_d8_DonorI.fastq.gz control_d8_DonorI.fastq.gz mv V_8_2_3_p600_d8_DonorII.fastq.gz control_d8_DonorII.fastq.gz mv V_8_4_2_p602_d8_DonorI.fastq.gz LT_d8_DonorI.fastq.gz mv V_8_4_1_p602_d8_DonorII.fastq.gz LT_d8_DonorII.fastq.gz mv V_8_2_4_p605_d8_DonorI.fastq.gz LTtr_d8_DonorI.fastq.gz mv V_8_2_3_p605_d8_DonorII.fastq.gz LTtr_d8_DonorII.fastq.gz -
Execute SPANDx to produce the mapping profile. For the GenBank file used, refer to the provided link.
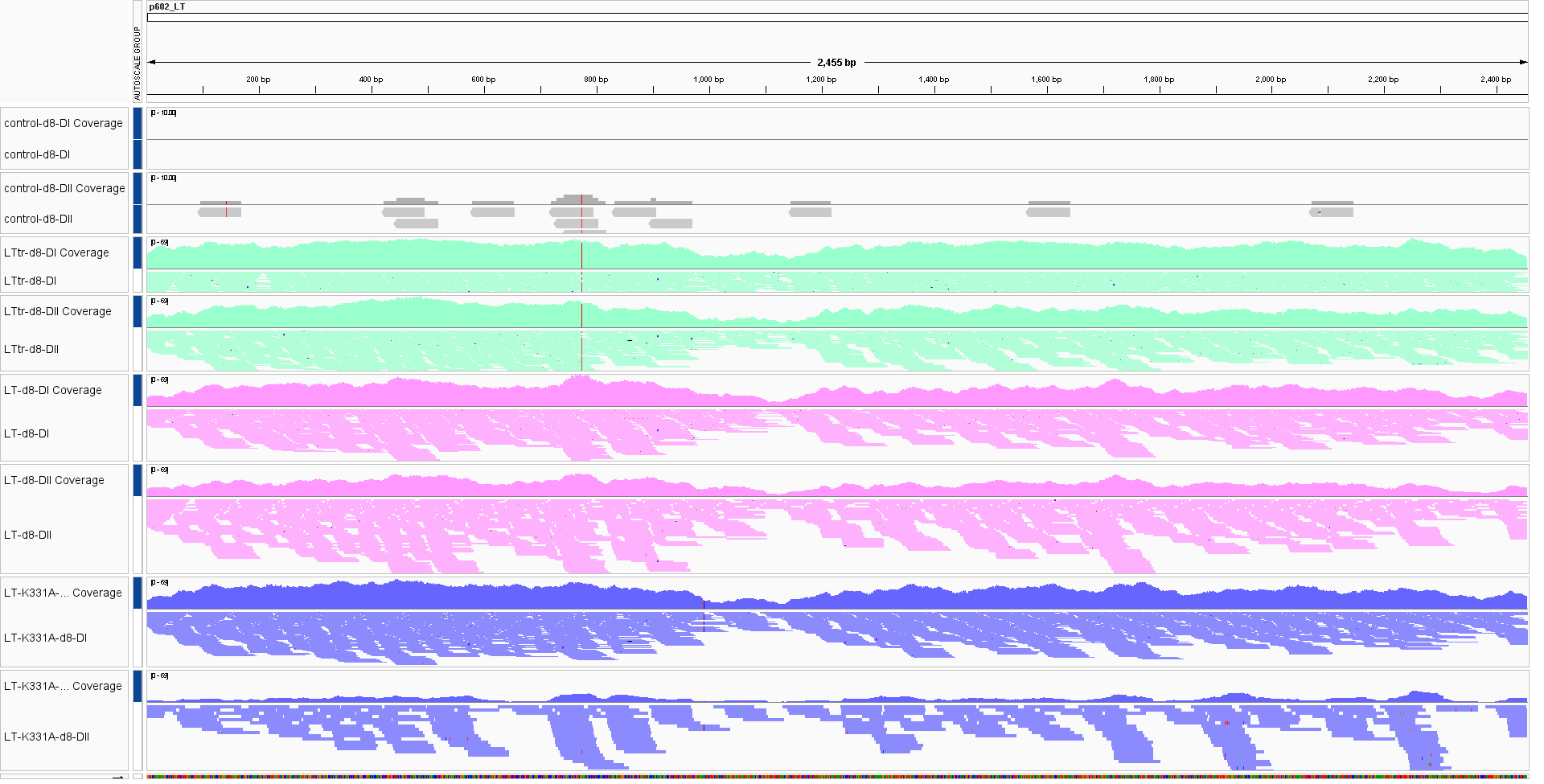
conda activate spandx [genbank copying] mkdir ~/anaconda3/envs/spandx/share/snpeff-4.3.1t-5/data/LT_wildtype cp LT_wt.gbk ~/anaconda3/envs/spandx/share/snpeff-4.3.1t-5/data/LT_wildtype/genes.gbk vim ~/anaconda3/envs/spandx/share/snpeff-4.3.1t-5/snpEff.config /home/jhuang/anaconda3/envs/spandx/bin/snpEff build -genbank LT_wildtype -d #The mutation in LTtr is located approximately at position 780, while the K331A mutation in LT is found near position 993 out of a total of 2454 nt. MDLVLNRKEREALCKLLEIAPNCYGNIPLMKAAFKRSCLKHHPDKGGNPVIMMELNTLWSKFQQNIHKLRSDFSMFDEVDEAPIYGTTKFKEWWRSGGFSFGKAYEYGPNPHGTNSRSRKPSSNASRGAPSGSSPPHSQSSSSGYGSFSASQASDSQSRGPDIPPEHHEEPTSSSGSSSREETTNSGRESSTPNGTSVPRNSSRTDGTWEDLFCDESLSSPEPPSSSEEPEEPPSSRSSPRQPPSSSAEEASSSQFTDEEYRSSSFTTPKTPPPFSRKRKFGGSRSSASSASSASFTSTPPKPKKNRETPVPTDFPIDLSDYLSHAVYSN<K>TVSCFAIYTTSDKAIELYDKIEKFKVDFKSRHACELGCILLFITLSKHRVSAIKNFCSTFCTISFLICKGVNKMPEMYNNLCKPPYKLLQENKPLLNYEFQEKEKEASCNWNLVAEFACEYELDDHFIILAHYLDFAKPFPCQKCENRSRLKPHKAHEAHHSNAKLFYESKSQKTICQQAADTVLAKRRLEMLEMTRTEMLCKKFKKHLERLRDLDTIDLLYYMGGVAWYCCLFEEFEKKLQKIIQLLTENIPKYRNIWFKGPINSGKTSFAAALIDLLEGKALNINCPSDKLPFELGCALDKFMVVFEDVKGQNSLNKDLQPGQGINNLDNLRDHLDGAVAVSLEKKHVNKKHQIFPPCIVTANDYFIPKTLIARFSYTLHFSPKANLRDSLDQNMEIRKRRILQSGTTLLLCLIWCLPDTTFKPCLQEEIKNWKQILQSEISYGKFCQMIENVEAGQDPLLNILIEEEGPEETEETQDSGTFSQ* nextflow run spandx/main.nf --fastq "Raw_Data_RNAseq_K331A_d8_SPANDx/*.fastq.gz" --ref LT_wt.fasta --annotation --database LT_wildtype --pairing SE -resume -
Open the BAM files created in the previous step using IGV.
点赞本文的读者
还没有人对此文章表态
本文有评论
没有评论
看文章,发评论,不要沉默
最受欢迎文章
- Motif Discovery in Biological Sequences: A Comparison of MEME and HOMER
- Calling peaks using findPeaks of HOMER
- Kraken2 Installation and Usage Guide
- Why Do Significant Gene Lists Change After Adding Additional Conditions in Differential Gene Expression Analysis?
- Should the inputs for GSVA be normalized or raw?
- PiCRUST2 Pipeline for Functional Prediction and Pathway Analysis in Metagenomics
- Updating Human Gene Identifiers using Ensembl BioMart: A Step-by-Step Guide
- pheatmap vs heatmap.2
- Setup conda environments
- Guide to Submitting Data to GEO (Gene Expression Omnibus)
最新文章
- Risks of Rebooting into Rescue Mode
- 足突(Podosome)、胞外囊泡(Extracellular Vesicle)与基质金属蛋白酶(MMPs)综合解析
- NCBI BioSample Submission Strategy for PJI and Nasal Microbiota Study
- Human RNA-seq processing for Data_Ben_RNAseq_2025
最多评论文章
- Updating Human Gene Identifiers using Ensembl BioMart: A Step-by-Step Guide
- The top 10 genes
- Retrieving KEGG Genes Using Bioservices in Python
推荐相似文章
All tools and services of BV-BRC
Enhanced Visualization of Gene Presence for the Selected Genes in Bongarts_S.epidermidis_HDRNA