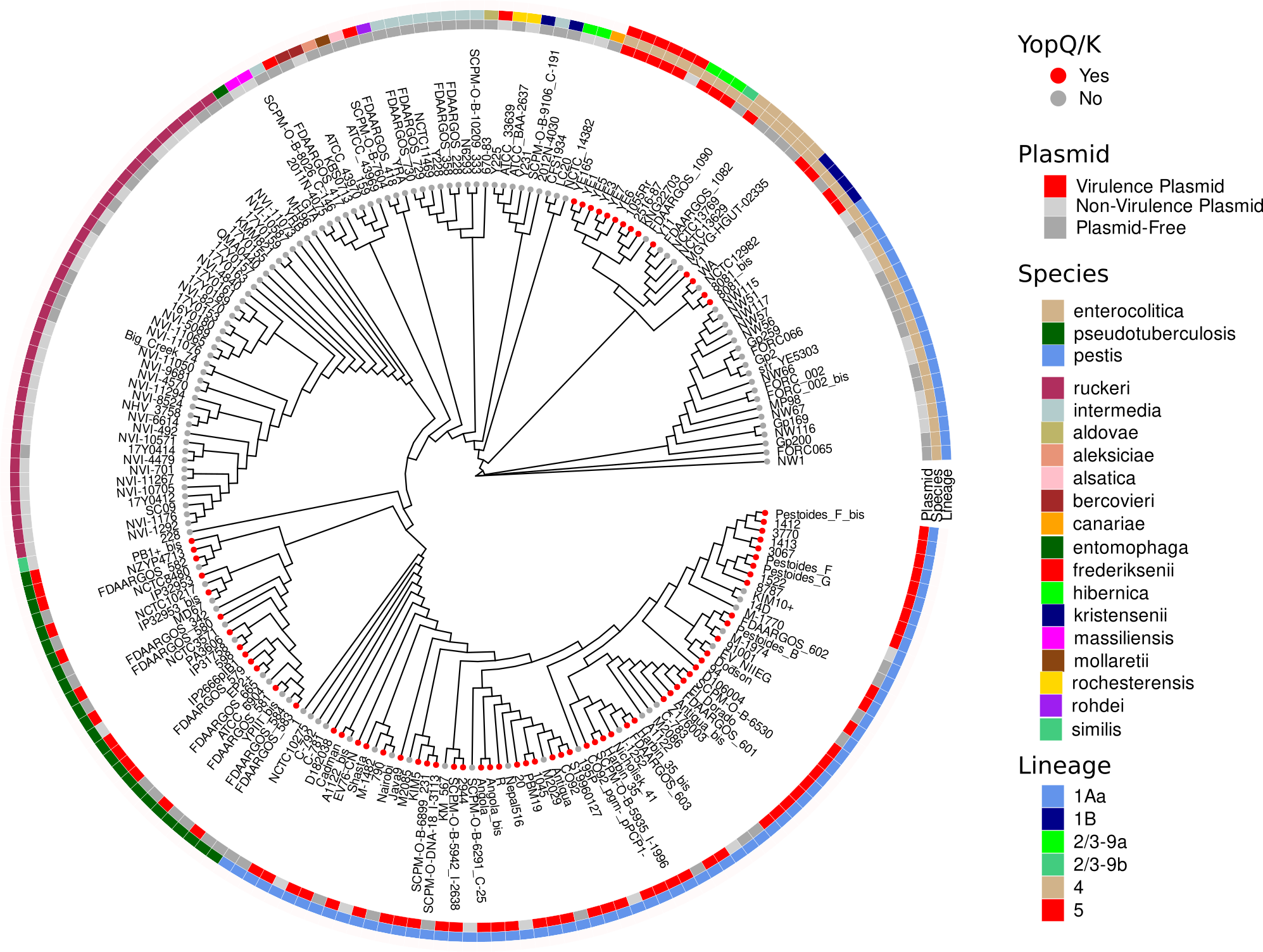
Virulence Plasmid Absence in Y. enterocolitica 1Aa Subcluster
gene_x 0 like s 832 view s
Tags: research
Pathogenicity of Yersinia enterocolitica biotype 1A https://academic.oup.com/femspd/article/38/2/127/531753
Abstract
- Yersinia enterocolitica strains of biotype 1A lack the known virulence determinants of strains in other categories, including the Yersinia virulence plasmid (pYV), and several chromosomal markers of pathogenicity.
- For this reason, and also because Y. enterocolitica strains of biotype 1A are frequently isolated from the environment or asymptomatic individuals, these bacteria are often assumed to be avirulent.
- On the other hand, there is a considerable body of clinical, epidemiological and experimental evidence to indicate that at least some strains of Y. enterocolitica biotype 1A are able to cause gastrointestinal symptoms which resemble those caused by pYV-bearing strains.
- The availability of a number of experimental systems, including cell culture and animal models of infection, provides an opportunity to identify and characterise the essential virulence determinants of biotype 1A strains.
-
Introduction
- The genus Yersinia encompasses a heterogeneous collection of facultatively anaerobic bacteria that belong to the family Enterobacteriaceae.
- Of the 11 species within this genus [1], only three, Y. pestis, Y. pseudotuberculosis and Y. enterocolitica, are regarded as pathogenic for humans.
- The virulence of bacteria in all three of these species correlates with carriage of a highly conserved plasmid, termed pYV [2], such that Yersinia spp. which lack this plasmid are considered avirulent.
- Of the three pYV-bearing species of Yersinia, Y. enterocolitica is the most heterogeneous, being divisible into approximately 30 distinct serotypes (on the basis of antigenic variation in cell wall lipopolysaccharide), and six biotypes (on the basis of variations in biochemical behaviour) (Tables 1 and 2).
- Of the six biotypes, biotype 1A is the most heterogeneous, and encompasses a wide range of serotypes (Table 2), of which serotypes O:5, O:6,30, O:6,31, O:7,8, O:10, as well as O-non-typable strains, are isolated most often (Table 3).
-
The diversity of biotype 1A strains is also evident when individual isolates are examined by techniques, such as ribotyping or pulsed-field gel electrophoresis, which show that strains of the same serotype, e.g. serotype O:5, show considerable genetic diversity, whereas pYV-bearing serotypes, including O:3 and O:9, are relatively well conserved [32,33].
-
One common feature of individual biotype 1A strains of Y. enterocolitica, however, is that they never carry pYV.
- For this reason, and also because biotype 1A strains are frequently isolated from the environment, strains of this biotype are often said to be avirulent.
- In this article, we review the epidemiological and experimental data regarding the pathogenicity of Y. enterocolitica biotype 1A and provide evidence that some strains of this biotype are able to cause disease by mechanisms that are independent of pYV or other known virulence determinants of Y. enterocolitica.
-
Virulence determinants of pYV-bearing strains of Y. enterocolitica
- Apart from pYV itself, pYV-bearing strains of Y. enterocolitica require a number of chromosomally encoded factors to express full virulence.
- Some of these factors are restricted to pYV-bearing bacteria, whereas others occur more widely. -[NOTE that translocation and secretion are different: translocation is yop from bacteria to host cell, secretion is yop from bacteria to environment.]
- Factors that are mostly limited to pYV-bearing strains of Y. enterocolitica include invasin, an outer membrane protein that is required for efficient translocation of bacteria across the intestinal epithelium [34]; Ail, another outer membrane protein that may contribute to adhesion, invasion and resistance to complement-mediated lysis [35]; Yst-a, a heat-stable enterotoxin that may contribute to the pathogenesis of diarrhoea associated with acute yersiniosis [36,37]; and Myf, a fimbrial antigen and putative adhesin [38].
- [NOTE that we can find the high-pathogenicity island by comparing 1A with 1B.]
- In addition, strains of biotype 1B, which are particularly virulent for humans and laboratory animals, carry a high-pathogenicity island which facilitates the uptake and utilisation of iron by bacterial cells, and hence may promote their growth under iron-limiting conditions in host tissues [39].
-
Virulence-associated determinants of pYV-bearing Y. enterocolitica that also occur in pYV-negative strains include cell surface lipopolysaccharide and SodA (a superoxide dismutase), which appear to facilitate bacterial survival in tissues [40,41], as well as urease, which enhances bacterial resistance to stomach acid and may also play a role in nitrogen assimilation [42,43].
-
pYV functions mainly as an anti-host genome that permits the bacteria which carry it to resist phagocytosis and complement-mediated lysis, thus allowing them to proliferate extracellularly in tissues.
- The genes carried on pYV include those for: (i) an outer membrane protein adhesin, YadA; (ii) a type III protein secretory apparatus which translocates effector proteins, known as Yops, from the bacterial cell to the cytoplasm of susceptible host cells; (iii) at least six distinct anti-host effector Yops; and (iv) factors involved in the regulation of Yop biosynthesis, secretion and translocation [2].
- The contribution of pYV-encoded factors, in particular YadA and the Yop effectors, to bacterial virulence has been established in a large number of experiments.
- Strains of yersiniae which lack pYV are susceptible to killing by complement and polymorphonuclear leukocytes, although they are able to persist in macrophages and non-professional phagocytic cells, and cause short-lived infections which typically are asymptomatic [44].
-
Evidence favouring the lack of virulence of biotype 1A strains (正)
- Biotype 1A strains of Y. enterocolitica are often considered to be non-pathogenic primarily because they do not possess the virulence-associated determinants of pYV-bearing strains.
- For example, pYV, which is considered indispensable for yersinia virulence, is never found in strains of biotype 1A [45].
- In addition, biotype 1A strains typically lack Ail, Myf, the ysa type III secretion system or the high-pathogenicity island, and only occasionally produce Yst-a [38,45^50].
- Another line of evidence that is taken to indicate the avirulence of biotype 1A strains of Y. enterocolitica is their relatively high prevalence in the environment and healthy animals.
- Indeed, biotype 1A strains are ubiquitous, inhabiting a wide variety of environmental niches such as soil and various sources of water, including streams, lakes, water wells, and wastewater [7,16,51^53].
- They are also frequently isolated from foods, including various vegetables and animal products, such as pork, poultry, packaged meat, seafoods, raw milk and pasteurised dairy products [16,54^60].
- Biotype 1A yersiniae are also found in a vast array of animals, including birds, fish, various insects, frogs and a wide range of mammals, including cattle, sheep, pigs and rodents [7,16,61].
- In most cases, animals that harbour biotype 1A strains are asymptomatic, thus giving support to the concept that these bacteria are avirulent commensals.
- There are also some epidemiological data to suggest that Y. enterocolitica biotype 1A strains are not pathogenic for humans.
-
For example, two large studies in Belgium, involving the microbiological investigation of more than 24 000 faecal samples over a period of almost 16 years, revealed that infection with biotype 1A yersiniae was not associated with gastrointestinal symptoms and that biotype 1A strains were more frequent amongst healthy subjects than in those who were being investigated for gastrointestinal complaints [62,63].
-
The relative avirulence of some biotype 1A isolates is supported by the results of several studies of experimental infection of animals.
- In contrast to pYV-bearing strains, which may cause a fatal infection in susceptible strains of mice, especially if the bacteria carry the high-pathogenicity island or the mice are pre-treated with iron or microbial siderophores [64,65], biotype 1A yersiniae are intrinsically avirulent for mice.
- Pai et al. [66] and Une [67] infected rabbits perorally with different biotype 1A strains from raw fish (serotype O:6,30) and pig intestine (serotype O:5), respectively, and concluded that these bacteria were avirulent.
- Similar findings were reported by Robins-Browne et al. [68] who found that gnotobiotic piglets, inoculated perorally with a biotype 1A strain of serotype O:5, which was originally isolated from milk, rapidly cleared the bacteria without developing any clinical or pathological evidence of disease.
- Interestingly, however, Corbel et al. [69] reported that a biotype 1A strain (serotype O:6) which was isolated from the liver of an aborted lamb, caused placentitis and abortion when inoculated into pregnant ewes.
- This finding suggests that animals may differ in their susceptibility to infection with biotype 1A yersiniae or that the pathogenicity of different biotype 1A strains may vary.
-
Evidence supporting the pathogenicity of biotype 1A strains of Y. enterocolitica (反)
- Despite the lack of identifiable virulence determinants in Y. enterocolitica strains of biotype 1A, these bacteria are frequently isolated from humans with gastrointestinal diseases as indicated in Table 3.
- Most of the studies listed in Table 3 are uncontrolled clinical observations, however, and therefore cannot be taken to indicate a causative association between Y. enterocolitica strains of biotype 1A and gastrointestinal complaints.
- In one study, however, Ebringer et al. [10] showed a significant association between infection with Y. enterocolitica biotype 1A and the occurrence of diarrhoea or other gastrointestinal symptoms.
- Moreover, in a prospective case-control study of infants with diarrhoea in Santiago, Chile, Morris et al. [28] found Y. enterocolitica in 1.9% of patients and 0.6% of age-matched controls (P = 0.05).
- Biotype 1A strains were isolated from seven children with diarrhoea, but not from any asymptomatic children (P = 0.02).
- Additional evidence of the clinical significance of biotype 1A yersiniae is that some patients infected with these bacteria show a speci¢c antibody response to the infecting strain [15,20].
- Although most reports of biotype 1A-associated disease refer to cases of sporadic infection, at least two outbreaks of gastrointestinal infection due to biotype 1A yersiniae have been reported.
- Ratnam et al. [18] in Canada described a nosocomial outbreak of diarrhoea that was attributed to a strain of biotype 1A, serogroup O:5, which affected nine hospitalised patients, and was evidently acquired from two patients who had been hospitalised for intermittent diarrhoea.
- More recently, Greenwood et al. [23] obtained Y. enterocolitica biotype 1A, serotype O:10, from 19 paediatric inpatients in a district general hospital in England over a 3-month period.
- Shortly afterwards, Y. enterocolitica biotype 1A, serotype O:6,30 was isolated from a further 17 hospitalised children in 1 month.
- The likely source of the O:6,30 strain was pasteurised milk that had been contaminated with a small quantity of raw milk [70].
- Published reports of infection with biotype 1A Y. enterocolitica are likely to under-represent the true prevalence of these infections, because many diagnostic laboratories will choose to disregard the potential clinical significance of these strains and not report them.
- In addition, laboratory techniques that are used to isolate Y. enterocolitica from faeces are not designed to isolate biotype 1A strains, and may even select against them.
- For example, because enrichment of cultures at 4°C is known to increase the likelihood of isolating biotype 1A Y. enterocolitica from faecal samples [8,14,18,19,62,71,72], some authorities have advised against the use of cold enrichment to reduce the chances of isolating ‘non-pathogenic’ strains [63,73].
-
Clinical manifestations of infections with biotype 1A Y. enterocolitica
- Infection with pYV-bearing strains of Y. enterocolitica may have protean manifestations depending on the age and individual susceptibility of the host.
- Among the more common presenting features are diarrhoea, especially in young children, enterocolitis, pseudo-appendicitis due to terminal ileitis, acute mesenteric lymphadenitis, pharyngitis and post-infection autoimmune sequelae, such as reactive arthritis and erythema nodosum.
- Infrequent, but potentially life-threatening complications include visceral abscesses, pneumonia, intussusception, endocarditis and septicaemia [74].
- By contrast, infections with biotype 1A strains are generally limited to the gastrointestinal tract with diarrhoea and abdominal pain the commonest symptoms.
- Nevertheless, some cases associated with reactive arthritis and other autoimmune complications more usually associated with pYV-positive strains of Y. enterocolitica have been described [10,20].
- Other symptoms of infection with biotype 1A yersiniae include fever, nausea, vomiting and malaise [17,18,25,28,75,76].
- Infection may also manifest as colitis, terminal ileitis and pseudo-appendicitis [17,76].
- A commonly reported feature of biotype 1A-associated diarrhoea is that the infection may persist for several weeks or months, sometimes involving cyclical periods of acute disease and remission [17-19,25,76].
- Infection with biotype 1A strains is said to be frequent in all age groups, in contrast to pYV-bearing strains which are more frequent in children [6,13,17,25].
- There is also evidence to suggest that infections with biotype 1A yersiniae are more common in individuals who are predisposed to infections in general [18,25], possibly attesting to the relatively low virulence of at least some biotype 1A strains.
- In general, however, acute infection with biotype 1A yersiniae tends to mimic infection with pYV-bearing Y. enterocolitica in terms of clinical manifestations and severity.
- A study by Burnens et al. [75] in Switzerland demonstrated that the duration and severity of infection associated with biotype 1A yersiniae was indistinguishable from that due to classical virulent biotypes.
-
Similar observations have been made in patients from England, Chile and Spain [26,28,77].
-
In contrast to pYV-bearing Y. enterocolitica, however, biotype 1A strains seldom give rise to extraintestinal infections or autoimmune sequelae.
- Nevertheless, biotype 1A strains have been isolated from abdominal exudates, wounds, sputum, urine, a labial ulcer and the gall bladder [7,12,19], and have been linked to cases of reactive arthritis [10].
- Presumably because they are susceptible to complement-mediated lysis, biotype 1A yersiniae rarely cause septicaemia and have only once been associated with a human death [21].
- In this case, a strain of serotype O:7,8 was isolated post mortem from the spleen and small intestine of a patient in Bangladesh, but Shigella boydii was also isolated from the large intestine of this patient [21].
- Patients who are convalescing from biotype 1A-induced gastroenteritis generally display low or undetectable titres of circulating antibodies to these bacteria [8].
- This has resulted in these strains being overlooked as aetiological agents, because pYV-bearing Y. enterocolitica, in particular serotype O:3, frequently evoke serum agglutinating antibody titres of greater than 1 in 200.
- However, low titres of antibody have also been found in patients with proven infection due to pYV-bearing Y. enterocolitica, including serotypes O:5,27, O:1,2,3 and O:21 [8,27], indicating that antibody titre does not necessarily correlate with the virulence of the infecting strain.
- Interestingly, Fletcher et al. [78] found that levels of secretory IgA antibodies in faeces directed against the patient’s own strain were similar in individuals infected with serotype O:3 yersiniae and biotype 1A strains.
-
Pathogenesis (6.1. Enterotoxins)
- Little is known of the pathogenic mechanisms of biotype 1A strains.
- Endoscopic examination of patients infected with these bacteria has revealed no obvious ulcerative or inflammatory changes [76], suggesting that the symptoms may be due to secreted toxins.
- When first isolated from clinical samples, most pYV-bearing strains of Y. enterocolitica secrete a low-molecular-mass, heat-stable enterotoxin, known as Yst-a [79].
- This toxin is active in infant mice, and may contribute to the production of diarrhoea by these bacteria [36].
- Although biotype 1A yersiniae seldom produce Yst-a, more than 80% of strains carry the ystB gene, which encodes a closely related, mouse-reactive toxin, known as Yst-b (Table 4) [50,80].
- The observation that many of the bacterial strains which carry the ystB gene do not produce detectable amounts of enterotoxin indicates that ystB is not always expressed in vitro.
- A similar situation pertains to pYV-bearing Y. enterocolitica, which, after repeated passage or prolonged storage, frequently lose the ability to produce Yst-a.
- This phenomenon is not caused by mutation of the ystA gene, but is due to silencing of this gene by YmoA (Yersinia modulator) [81].
- As biotype 1A yersiniae also carry the ymoA gene (unpublished data), it is conceivable that in these bacteria YmoA is able to silence ystB.
- However, it is also possible that ystB is non-functional in some biotype 1A strains.
- One biotype 1A strain of serogroup O:6 that was isolated from a child with diarrhoea, secretes a novel heat-stable enterotoxin termed YST II, which is also mouse-reactive, but has a distinct mechanism of action compared with other heat-stable enterotoxins [82].
- The gene encoding YST II has not been identi¢ed, and hence the prevalence of YST II production amongst biotype 1A strains in general has not been elucidated.
-
Pathogenesis (6.2. Fimbrial adhesins)
- Another potential virulence factor of biotype 1A yersiniae are ¢mbriae, which take various forms in these bacteria.
- One of these, designated MR/Y-HA, is 8 nm in diameter, agglutinates erythrocytes of 10 di¡erent animal species in the presence of mannose, and is expressed in vitro at low temperatures, but not at 37°C [83].
- A second type of ¢mbria, designated MR/K-like HA, is 4 nm in diameter and mediates mannose-resistant haemagglutination of chicken erythrocytes, but not erythrocytes from a variety of other species [83].
- Expression of these fimbriae in vitro occurs only after serial passaging of bacteria for at least 7 days.
- Moreover, as they do not mediate adherence of bacteria to cultured epithelial cells [84], their contribution to the pathogenesis of infection with biotype 1A strains is unknown.
- Some strains of Y. enterocolitica produce a ¢mbrial adhesin, named Myf (for mucoid Yersinia fibrillae), because it bestows a mucoid appearance on bacterial colonies which express it [38].
- Myf are narrow flexible fimbriae which resemble CS3, an essential colonisation factor of some human strains of enterotoxigenic Escherichia coli [85].
- MyfA, the major structural subunit of Myf, shows some homology to the PapG protein of pyelonephritis-associated strains of E. coli, and is 44% identical at the DNA level to the pH6 antigen of Y. pseudotuberculosis and Y. pestis, which also has a ¢brillar structure and mediates thermoinducible binding of Y. pseudotuberculosis to tissue culture cells [86,87].
- As with Yst-a, however, myf genes occur in a minority of biotype 1A strains [50] (Table 4), and their contribution to the virulence of these bacteria is unknown.
-
Pathogenesis (6.3. Interaction with epithelial cells and macrophages)
- Preliminary observations in our laboratory indicated that a biotype 1A isolate, which was obtained from a child with diarrhoea [28,82], was able to resist killing by murine peritoneal macrophages in vitro.
- We also showed that this was not a uniform characteristic of biotype 1A Y. enterocolitica, as another serotype O:6 strain, which was originally isolated from milk [68], was killed by macrophages under the same experimental conditions.
- As biotype 1A strains of Y. enterocolitica comprise a heterogeneous group of bacteria which occupy a wide range of environmental niches, we postulated there may be a pathogenic subgroup of these bacteria which cannot be readily identified because they lack the known virulence determinants of pYV-bearing strains.
- To learn more about the possible virulence mechanisms of biotype 1A yersiniae, we divided the biotype 1A strains in our culture collection into two groups.
- One group comprised strains isolated from patients with gastrointestinal symptoms consistent with yersiniosis, and therefore was hypothesised to include mostly virulent strains.
- The second group comprised strains isolated from environmental sources, which were postulated to be saprophytic, and incapable of causing disease.
- Initially, we examined the two groups of bacteria for their carriage of known or suspected virulence-associated genes of Y. enterocolitica, including ail, myfA, ystA, ystB and virulence genes borne on pYV [50].
- The results of this investigation showed that apart from ystB, which was detected in more than 80% of strains, fewer than 15% of biotype 1A strains carried sequences that were homologous to virulence-associated genes of pYV-bearing bacteria (Table 4).
- We also found that the frequency of these genes was similar in strains of clinical and environmental origin.
- The ability of Y. enterocolitica to invade epithelial cells is an important correlate of virulence [46,91].
- One reason that strains of biotype 1A have been considered to be avirulent is that they invade tissue culture cells to a lesser extent than pYV-bearing strains [88,89], although, paradoxically, pYV itself may retard cell invasion via the effects of translocated Yops on cytoskeletal proteins [2].
-
We have confirmed that biotype 1A strains penetrate HEp-2 cells, a continuous line of human-derived epithelial cells, less effectively than pYV-cured strains [50]. Interestingly, however, we observed that strains of clinical origin invaded HEp-2 cells, Chinese hamster ovary cells and T84 cells, a polarised epithelial cell line of intestinal origin, to a significantly greater extent than strains obtained from other sources (Fig. 1) [50].
-
Electron microscopic examination of HEp-2 and bone marrow-derived macrophages incubated with clinical isolates of Y. enterocolitica biotype 1A revealed a novel pattern of invasion, in that some cells contained large numbers of internalised bacteria which were sametimes located within vacuoles, but most cells were spared (Figs. 2A, 3A and 3B).
- By contrast, cells infected with biotype 1A strains of environmental origin contained no more than two (often degraded) bacteria per cell, with the great majority of cells being uninfected (Fig. 2B and 3C).
- Clinical isolates of Y. enterocolitica strains of biotypes 1B and 2, which bore classical chromosomally encoded virulence markers, but were cured of pYV, invaded cells in a more even distribution than that observed with biotype 1A yersiniae (Fig. 2C).
- These findings suggest that biotype 1A strains of Y. enterocolitica may invade cells by a novel mechanism which differs from that employed by pYV-bearing strains.
-
In keeping with our original observation that a biotype 1A strain of clinical origin was more resistant to killing by macrophages than an environmental isolate, we subsequently showed that relative resistance to macrophage-induced killing is a characteristic of clinical biotype 1A strains in general, although there is considerable overlap between individual clinical and environmental strains [90].
-
Interestingly, we also found that clinical strains are able to replicate within epithelial cells and macrophages and then escape from these cells, whereas environmental strains are not [90].
- The mechanism of bacterial escape is unknown, but insofar as it does not involve host cell lysis, it appears to resemble exocytosis [90].
- Recently, we have used transposon mutagenesis in an attempt to identify the bacterial genes required for escape.
- Preliminary results of this work indicate that bacteria require intact cell wall lipopolysaccharide to replicate within host cells before they can escape.
- We have also used genomic subtractive hybridisation to determine genetic differences between a clinical and an environmental strain of Y. enterocolitica biotype 1A, with the aim of identifying putative virulence-associated genes in the clinical strain [91].
- This strategy yielded 54 DNA fragments that were speci¢c for the clinical strain. Of these, 33 sequences had signi¢cant matches with known proteins in the GenBank database, such as a flagellar hook-associated protein, an adenine methyltransferase and an autotransporter protein.
- Interestingly, four fragments exhibited homology to three insecticidal toxin complex (tc) genes that were ¢rst identi¢ed in Photorhabdus luminescens [92] and are also present in Y. pestis [93,94].
- The contribution of these genes to the virulence of Y. enterocolitica is not known.
-
Pathogenesis (6.4. Animal models of infection)
- Studies to identify the essential virulence determinants of pYV-bearing Y. enterocolitica have been facilitated by access to animal models of infection, such as guinea pigs or mice pre-treated with iron and desferrioxamine [65,95].
- The latter is a microbial siderophore that permits pYV-bearing strains of Y. enterocolitica, which lack the high-pathogenicity island [39], to grow under iron-limiting conditions in vitro and in vivo [65].
- Until recently, studies to identify virulence determinants of biotype 1A yersiniae were hampered by the lack of a suitable animal model.
- We have shown, however, that mice inoculated perorally with 10 9 colony-forming units of biotype 1A Y. enterocolitica may excrete the bacteria for up to 10 days (Fig. 4).
- Importantly, strains of clinical origin colonise both the small and large intestine of mice for signi¢cantly longer periods than strains obtained from water or milk [50].
- The extent and duration of bacterial excretion in the faeces in this model were enhanced somewhat by pre-treating the mice with iron and desferriox-amine.
-
Summary and conclusions
- Despite the absence of recognised virulence determinants in biotype 1A strains of Y. enterocolitica, there is considerable clinical, epidemiological and experimental evidence to indicate that at least some of these strains are able to cause gastrointestinal disease in humans that clinically resembles yersiniosis caused by pYV-bearing Y. enterocolitica strains of other biotypes. In this respect, Y. enterocolitica may resemble enterovirulent strains of E. coli, which are classified into distinct pathotypes that differ from each other in terms of their specific virulence determinants [96].
- Our observations that biotype 1A strains of clinical origin di¡er signi¢cantly from non-clinical isolates in terms of their greater capacity to (1) penetrate cultured epithelial cells, (2) survive within macrophages, (3) exocytose from epithelial cells and macrophages and (4) colonise the intestinal tract of orally inoculated mice for prolonged periods provide a variety of convenient models in which to assay the pathogenicity of these bacteria.
- These models could be used to identify and characterise the virulence determinants of biotype 1A yersiniae more precisely.
- Once these factors are identified, it will be important to determine if they also contribute to the virulence of pYV-bearing strains of Y. enterocolitica, as well as pYV-negative strains of biotypes other than 1A (Table 2), and even other Yersinia species, such as Y. frederiksenii and Y. bercovieri, which have been associated with symptomatic infections in humans, and which also lack pYV [97].
点赞本文的读者
还没有人对此文章表态
本文有评论
没有评论
看文章,发评论,不要沉默
最受欢迎文章
- Motif Discovery in Biological Sequences: A Comparison of MEME and HOMER
- Why Do Significant Gene Lists Change After Adding Additional Conditions in Differential Gene Expression Analysis?
- Calling peaks using findPeaks of HOMER
- Updating Human Gene Identifiers using Ensembl BioMart: A Step-by-Step Guide
- pheatmap vs heatmap.2
- Should the inputs for GSVA be normalized or raw?
- Setup conda environments
- PiCRUST2 Pipeline for Functional Prediction and Pathway Analysis in Metagenomics
- Kraken2 Installation and Usage Guide
- File format for single channel analysis of Agilent microarray data with Limma?
最新文章
- Workflow using PICRUSt2 for Data_Karoline_16S_2025
- Viral genome assembly and recombination analysis for Data_Sophie_HDV_Sequences
- 阳光房漏水怎么办?丁基胶带才是最佳密封选择
- DAMIAN Post-processing for Flavivirus and FSME
最多评论文章
- Updating Human Gene Identifiers using Ensembl BioMart: A Step-by-Step Guide
- The top 10 genes
- Retrieving KEGG Genes Using Bioservices in Python
推荐相似文章
Distinguishing Cis and Trans Regulation: A Focus on miRNAs
Beitragsbemessungsgrenzen und Rechengrößen in der Sozialversicherung 2023